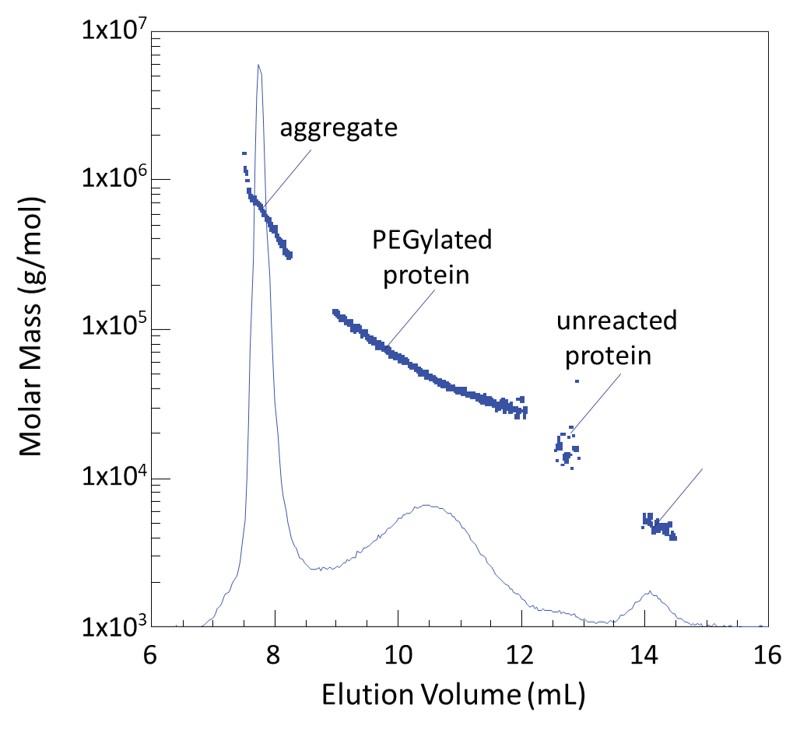
A protocol for recombinant protein quantification by densitometry - Alonso Villela - 2020 - MicrobiologyOpen - Wiley Online Library
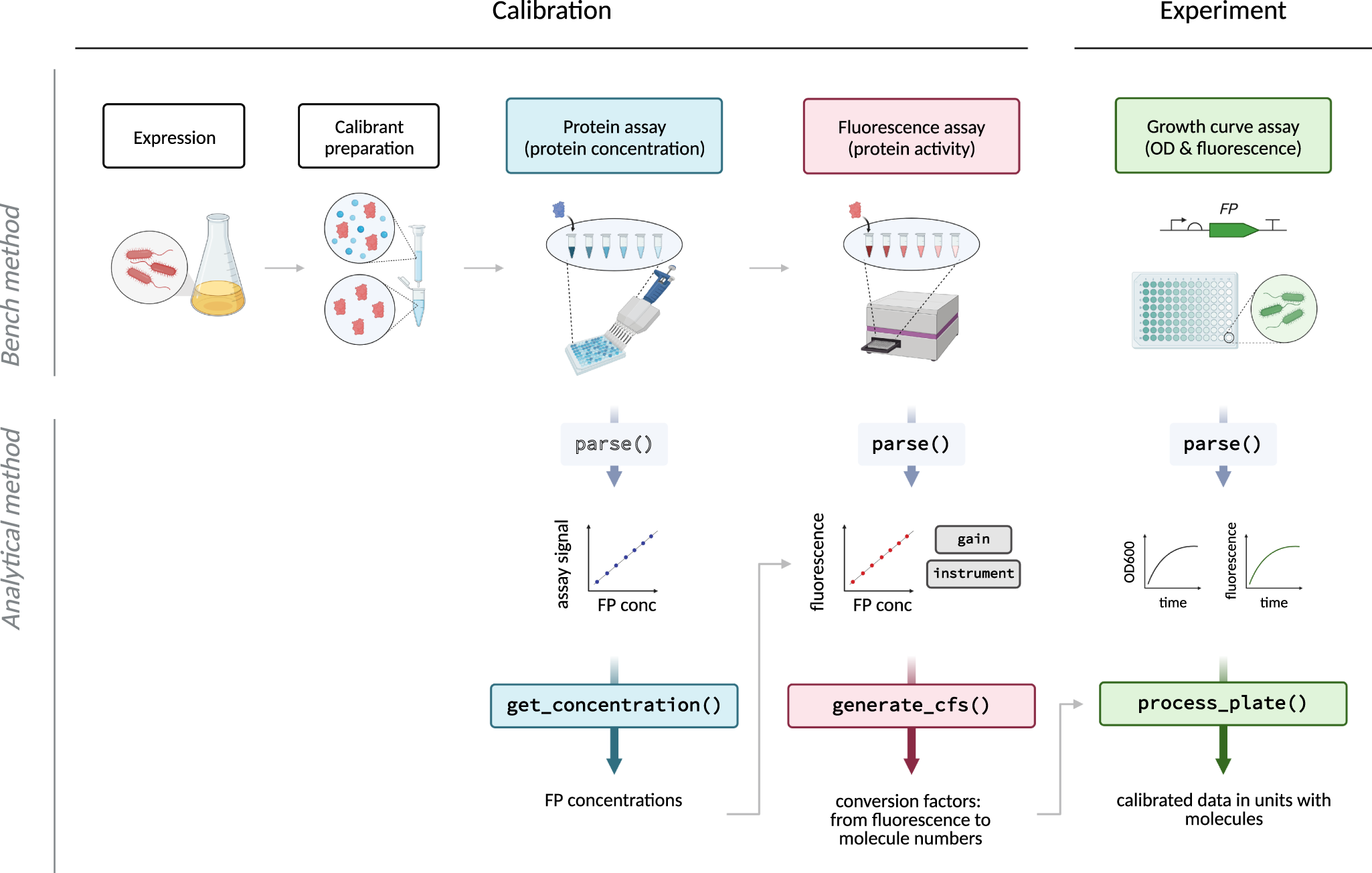
Absolute protein quantification using fluorescence measurements with FPCountR | Nature Communications
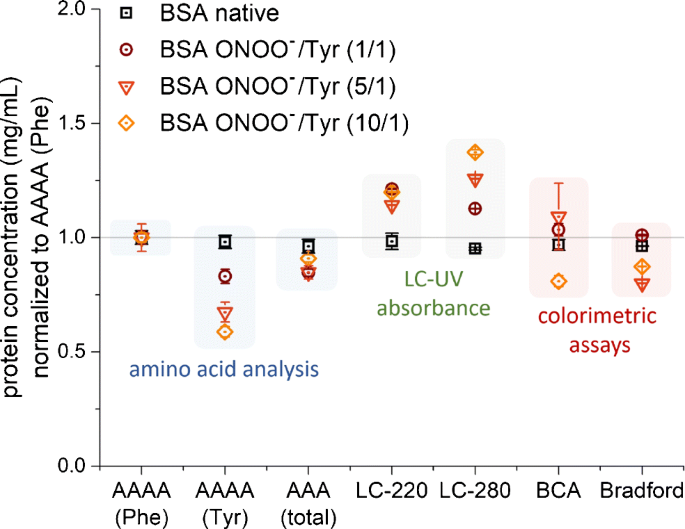
Determination of the protein content of complex samples by aromatic amino acid analysis, liquid chromatography-UV absorbance, and colorimetry | SpringerLink
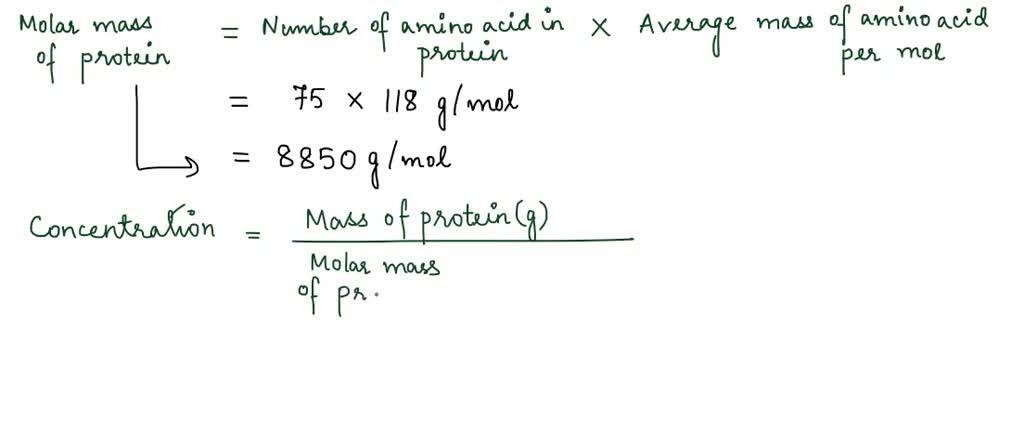
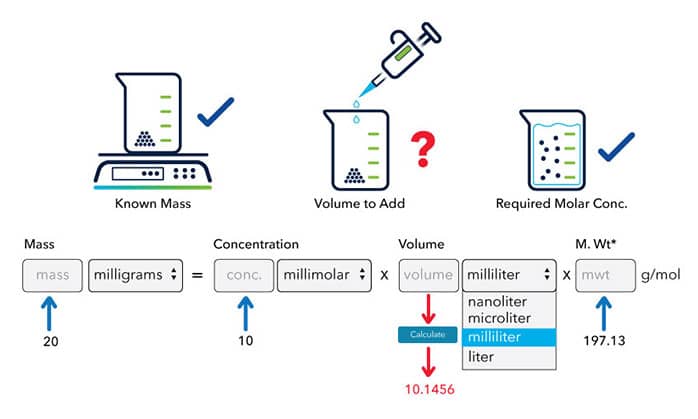
![Molarity Calculator [with Molar Formula] Molarity Calculator [with Molar Formula]](https://scrn-cdn.omnicalculator.com/chemistry/molarity@2.png)




.png?revision=1&size=bestfit&width=367&height=304)